Parallel Pipeline (Hyperparameter Search)
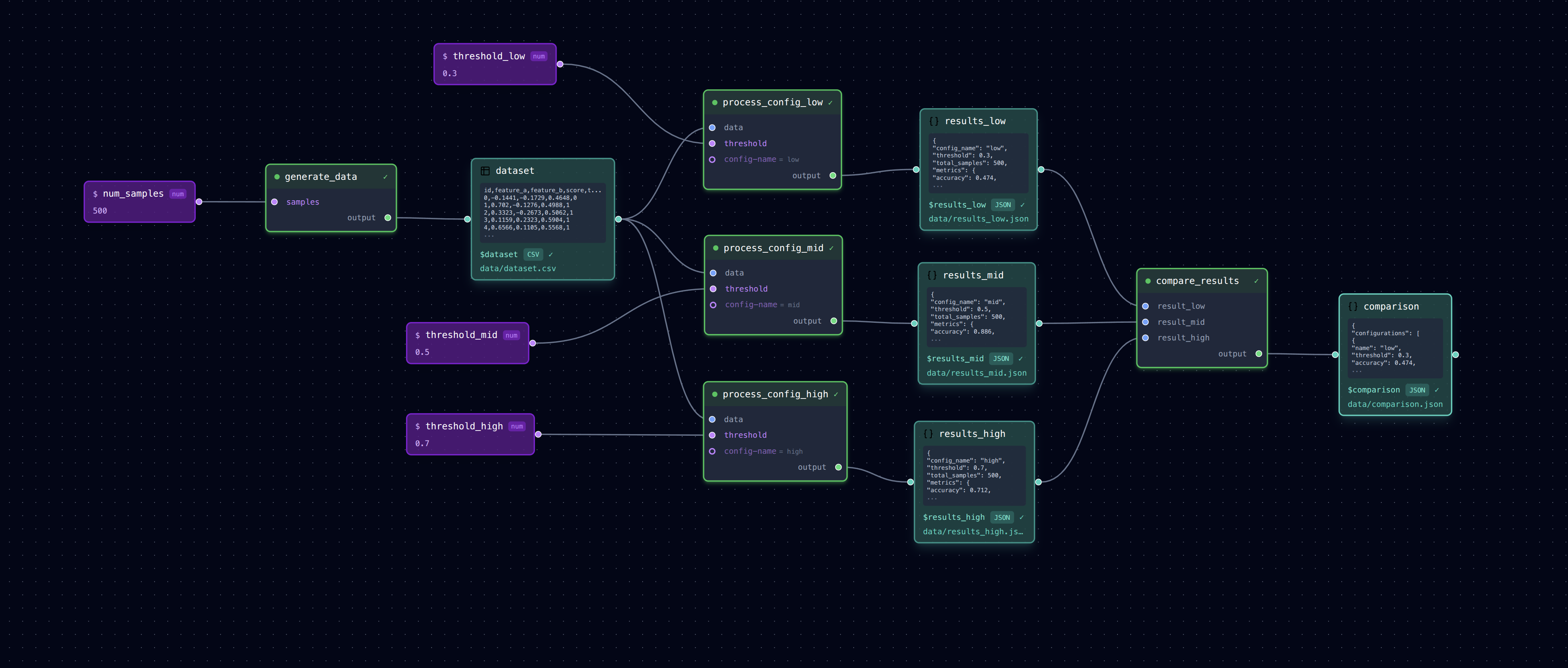
Running the same processing with different parameter configurations to compare results.
┌─► process (config A) ─► results_a.json ──┐
generate_data ──────┼─► process (config B) ─► results_b.json ──┼─► compare_results
└─► process (config C) ─► results_c.json ──┘
What It Does
This pattern is common in research: generate data once, then run the same analysis with different hyperparameters to find the best configuration.
- generate_data: Creates synthetic classification data
- process_config_a/b/c: Three parallel runs with different thresholds
- compare_results: Aggregates all results into a comparison table
Each "process" step uses a different threshold parameter, producing separate outputs.
Run It
# Run full pipeline (parallel execution enabled - configs run concurrently)
loom examples/parallel/pipeline.yml
# Run sequentially instead
loom examples/parallel/pipeline.yml --sequential
# Check outputs
cat examples/parallel/data/dataset.csv # Generated data
cat examples/parallel/data/results_low.json # Threshold 0.3
cat examples/parallel/data/results_mid.json # Threshold 0.5
cat examples/parallel/data/results_high.json # Threshold 0.7
cat examples/parallel/data/comparison.json # Side-by-side comparison
# Run just one configuration branch
loom examples/parallel/pipeline.yml --step process_config_low
# Open in editor to see the parallel structure
loom-ui examples/parallel/pipeline.yml
Parallel Execution
This pipeline uses parallel execution (configured in pipeline.yml):
With this configuration, the three process_config_* steps run concurrently after generate_data completes. The compare_results step waits for all three to finish before running.
Note: The process.py task includes a random 2-5 second delay to make parallel execution visible. When running in parallel, all three process steps start together and complete in ~5 seconds total. Running sequentially (--sequential) takes ~12 seconds since each step waits for the previous one.
Pattern: Hyperparameter Search
The key pattern here is: 1. One input source feeds multiple processing branches 2. Each branch has different parameters but the same task script 3. Results are aggregated at the end for comparison
This is useful for: - Trying different model hyperparameters - Comparing algorithm variants - A/B testing processing approaches
Extending This Pattern
To add more configurations:
1. Add a new variable for the output: results_d: data/results_d.json
2. Copy one of the process_config_* steps, rename it, change the threshold
3. Add the new result to compare_results inputs
Files
pipeline.yml— Pipeline with parallel branchestasks/generate_data.py— Creates synthetic datatasks/process.py— Classifies data with configurable thresholdtasks/compare_results.py— Aggregates results for comparison